Here is a guide for the kind of person who needs to get their data into R and have never done so or are struggling to get their data to load. I’ve tried to explain using simple words and lots of detail – the post is aimed at people who are not comfortable with code, so programmers might find it too simplified!
So, let’s start with the basics. Why are we bothering? At the beginning, we bother for three mean reasons, IMHO.
- When you write some code, you’ve got a record of exactly what you’ve done to your data. This means that the methods section is much easier to write.
- This also means that if you have to change one little annoying thing in your data file, you can run all the analyses again on one mouse click. For me, this is the most important reason!
- You have much better control over settings, particularly in graphics. That way, you don’t end up with a horrible excel-default-settings figure (which a journal will reject).
Of course, advanced users will also highlight that R has any function you can dream of, or at least the building blocks to make that function. And best of all R is free!
Before you begin, you really really should use a text editor for writing your code, not the R GUI (graphical user interface – the windows which open when you select R in your list of programs). This is because text editors do useful things like save your code, and colour it in pretty colours for the names of functions and arguments – the same as the code in this blog. Just believe me, this is extremely useful for finding mistakes. I use Tinn-R but nearly everyone is now using R Studio – both are free and open-source. Find a text editor that works with R and use it!
Now, R thinks about the world a particular way. R divides the world into objects, functions and arguments. R commands are very often in the form “Make me an object that is made by doing these functions to that other object, according to those arguments”. Arguments are kind of like settings. Your data, as far as R sees it, is an object. But it’s probably an Excel or Open Office spreadsheet, which R does not deal directly with and cannot use as an object. R would much rather that your data was a data object, so you want to make a data object. Many first-timers use the clipboard function, and we often do that in teaching because we have the datasets ready and in the right format. Chances are that you don’t have your data in the right format, so we’re going to take a slightly longer route which we have much more control over.
I’m going to describe how to load a “fat” (variables x samples) table. Examples might be quadrats and plant species, or water samples and chemical measurements (pH, salinity, temperature). You should prepare the Excel file with the variables along the top row, and the site or sample number down the first column. There should not be any special characters (stuff like / ! #) though Norwegian letters should be ok. Variable names need to start with a letter – microbiologists using numbers to identify genes should just add an “a” in front, it can be taken out later if needed. Numbers are fine for sample names (as are number-letter combinations). Save the spreadsheet as a tab-delimited text file. Here’s the top corner of an excel spreadsheet set up correctly:
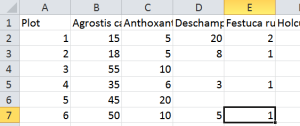
Next, open your text editor. The first thing to do is write a line of code to tell R where you keep your data. Then R will always look there to open files, and it will always put figures and so on in the same place. This place is called the ‘working directory’. So on my computer at home, I do this:
setwd("\\\\C:\\MyDocuments\\ Rstuff")
Rstuff is the folder where I’m storing that text file of data, and where I want the graphs to end up. See all those \\ where the file path normally just has the one \? They are because R sees \ as a ‘special character’, so you need to tell it “no R, that really is a file directory marker, not a special thingy.” The absolute easiest way to do this is with something called file.choose, which opens a regular window and lets you browse to any file in the folder that you want. That puts the file path in the r window in the right format, and you can copy the folders bit of that file path to the setwd function.
file.choose()
The brackets are important, but leave them empty like I’ve shown.
Now we make a data object in R by using the read.table function. I’ve made an example of data that you can download by clicking: community_data,
but you should try this with your own dataset. We made our Excel spreadsheet into a .txt file, because it makes this next step a lot simpler and more reliable.
community.df<-read.table ("Community_data.txt", sep="\t", header=TRUE, row.names=1)
The .df means data frame, a kind of data object. Putting that in the new object’s name doesn’t force R to make it a data frame, but helps you remember what you were intending. The <- (an arrow made out of a “less than” sign and a hyphen) tells R that on the left of the arrow is what you want it to make, and on the right is how you want it made. The “read.table” is the function – what it is you are asking R to do. "Community_data.txt" is the name of the data file – don’t forget the "…" and the .txt at the end. The “sep=\t” tells R that this is a tab delimited file. The “header=TRUE” tells R that your data has a header, that is, that the first row of the .txt file is the column names rather than data. Finally, “row.names=1” tells R that you have sample numbers in the first column.
So now we have two lines of code in our text editor. Most text editors will send code to R by clicking a button. I’m going to let the text editor help files tell you how to do that, because it varies between text editors and operating systems. So now send those two lines.
Did it work? Let’s look at the data. We can ask R what it thinks the data object is.
str(community.df)
This command does not change your data, but just displays the structure of the data. You can check that the file has read in correctly, has the right number of species (columns) and samples (rows). The example data should say “data.frame': 10 obs. of 11 variables:” and then list the species names.
You can look at all the data at once, by just typing the name of the data object
community.df
But this is a bad idea if you have more than ten samples. Instead
head(community.df)
shows you the first six rows.
Here is an example of how it should look.
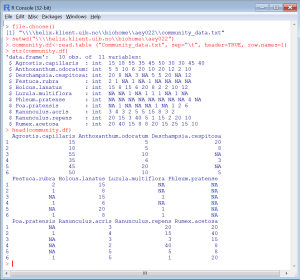
Not working?
A common error message comes at the read.table stage.
Error in file(file, "rt") : cannot open the connection In addition: Warning message: In file(file, "rt") : cannot open file 'Communitydata.txt': No such file or directory
This means that either you have set the working directory wrong, or typed the filename wrong. Remember to type the suffix for the file type, here .txt!
Well explained!